Description
New insight gained on breast cancer progression, treatment

Vimla Band, PhD
A new protein biomarker discovered by University of Nebraska Medical Center researchers could help in the prognosis and treatment of breast cancer.
Called ADA3, the biomarker could remove a major blind spot for physicians treating patients with breast cancer. Accurately classifying cancer and then determining the best course of treatment has been a major obstacle, but ADA3 opens a window to new insights about breast cancer progression.
When ADA3 is found in a cancer cell’s nucleus, patients typically have a more promising prognosis with tumors that grow slower and are less invasive. However, when ADA3 is found in the cytoplasm—the goo between the cell wall and nucleus—patients face a far more aggressive type of breast cancer, and a less promising prognosis.
Researchers also found that ADA3 correlates with two important breast cancer indicators: Estrogen receptor, ER for short, and human epidermal growth factor receptor 2 or HER2.
With further development, ADA3 could not only provide future physicians greater insight into how a patient’s breast cancer might progress, but how well that patient will respond to various treatment options such as so-called anti-ER and anti-HER2 drugs.
To discuss licensing opportunities please contact Matt Boehm, PhD at mboehm@unmc.edu or 402-536-9881.
Technical Data
Breast cancer biomarker for the prediction of disease progression and survival
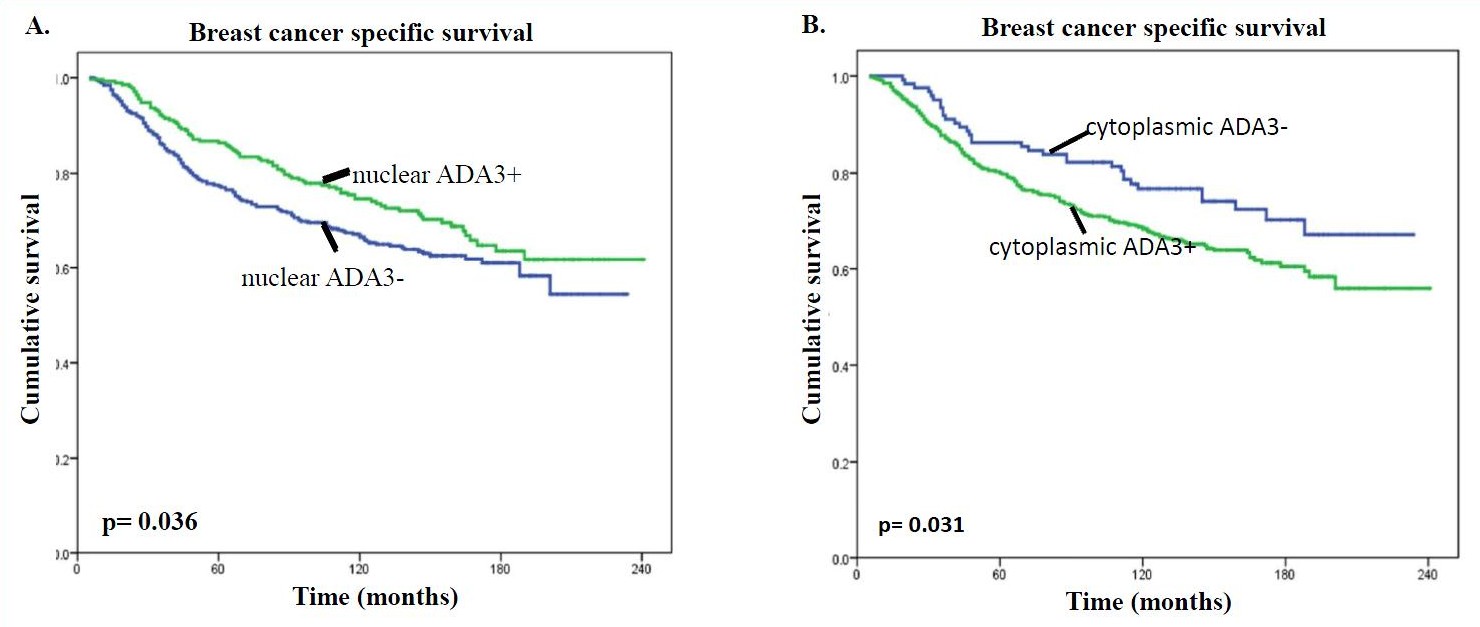
Researchers at the University of Nebraska Medical Center have identified a novel protein biomarker, known as ADA3, useful for assessing the prognosis of breast cancer patients. ADA3 expression and subcellular localization were analyzed through immunohistochemistry in 900 normal and breast cancer tissue specimens. Nuclear localization of ADA3 in breast cancer specimens was associated with a more differentiated phenotype (Grade 1 or 2), a lower proliferative index, an absence of vascular invasion, and a more favorable (excellent to good) Nottingham Prognostic Index (NPI). On the other hand, cytoplasmic localization of ADA3 was associated with more aggressive morphological and molecular features including a poorly differentiated phenotype (Grade 3), a high proliferation status, positive vascular invasion, and a less favorable (poor) NPI (Fig. A & B).
 Comparison of ADA3 expression with specific breast cancer subtypes revealed a strong positive association between nuclear ADA3 staining and Estrogen Receptor (ER) and Progesterone Receptor (PR) status and of ER-positive luminal breast cancers in general. Predominantly cytoplasmic ADA3 staining, on the other hand, revealed a negative association with ER and PR status; however, was positively associated with the receptor tyrosine kinases, HER2 and EGFR.
Comparison of ADA3 expression with specific breast cancer subtypes revealed a strong positive association between nuclear ADA3 staining and Estrogen Receptor (ER) and Progesterone Receptor (PR) status and of ER-positive luminal breast cancers in general. Predominantly cytoplasmic ADA3 staining, on the other hand, revealed a negative association with ER and PR status; however, was positively associated with the receptor tyrosine kinases, HER2 and EGFR.
It was also found that predominantly cytoplasmic ADA3 in ER-negative tumors was associated with a worse clinical outcome than patients lacking cytoplasmic ADA3 in ER-positive tumors (Fig. E & F).
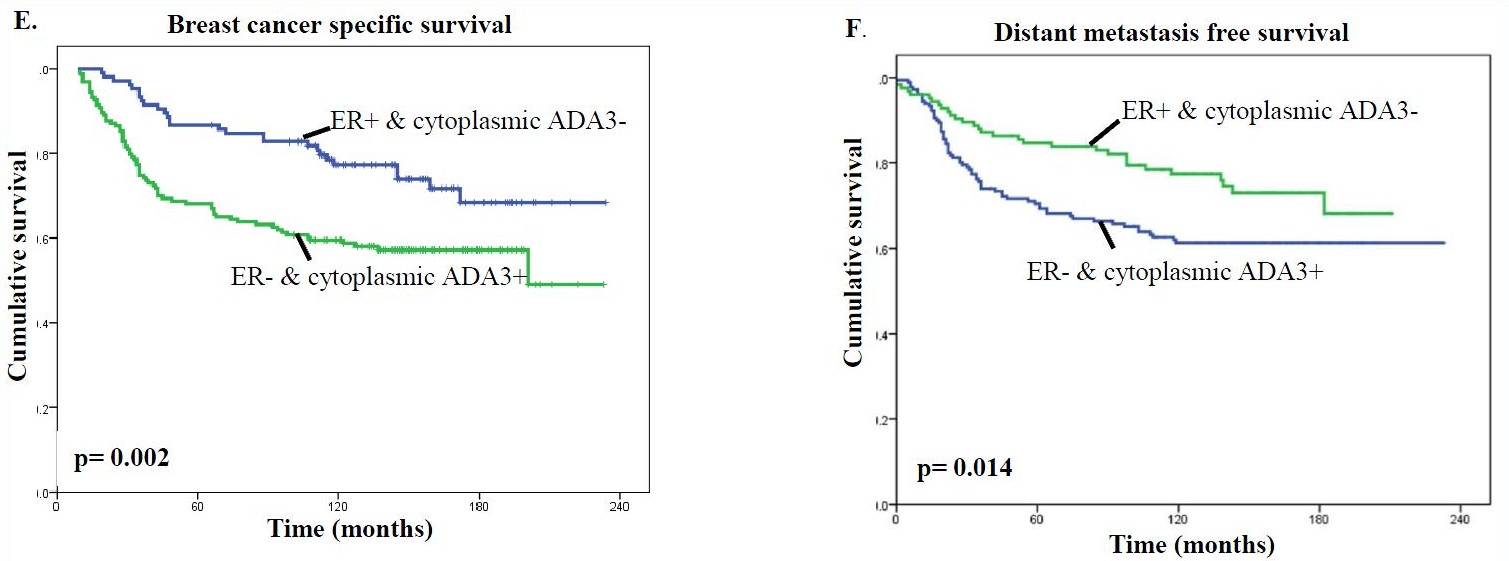
In addition, the presence of high levels of cytoplasmic ADA3 in HER2 or EGFR positive tumors was associated with significantly worse patient survival compared with samples lacking HER/EGFR expression and even more so compared with samples lacking HER2/EGFR expression and cytoplasmic ADA3 for both BCSS (Fig. G & H) and DMFS (Fig. I & J) analyses.
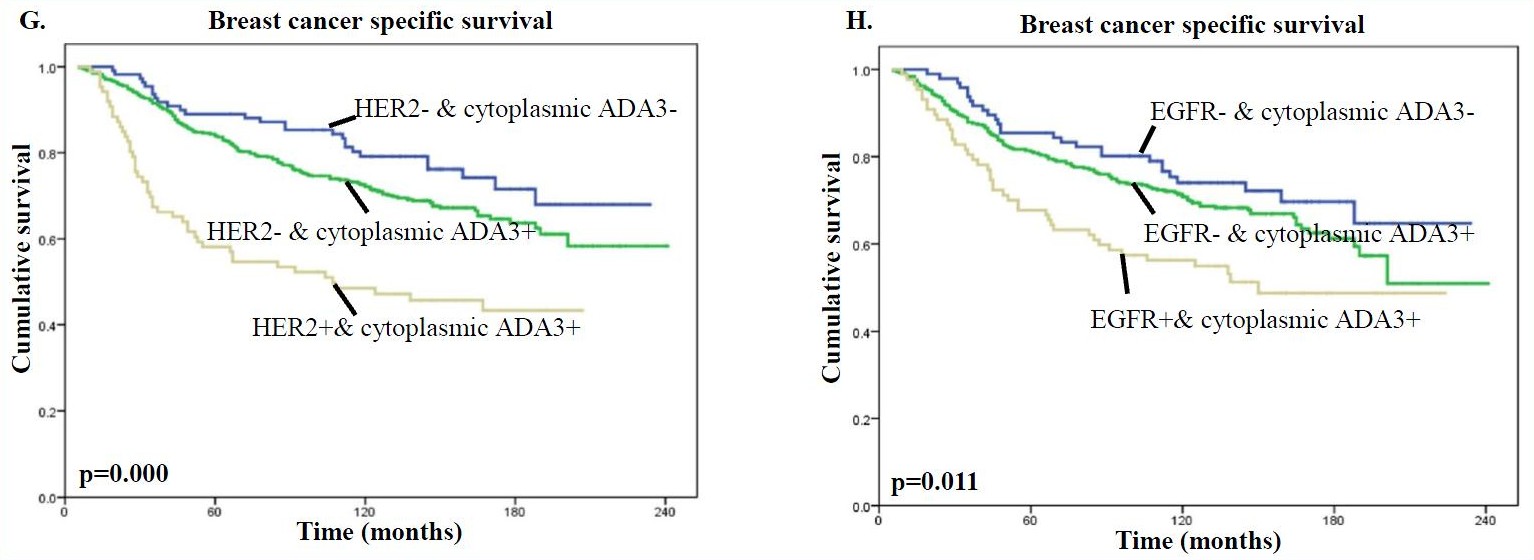
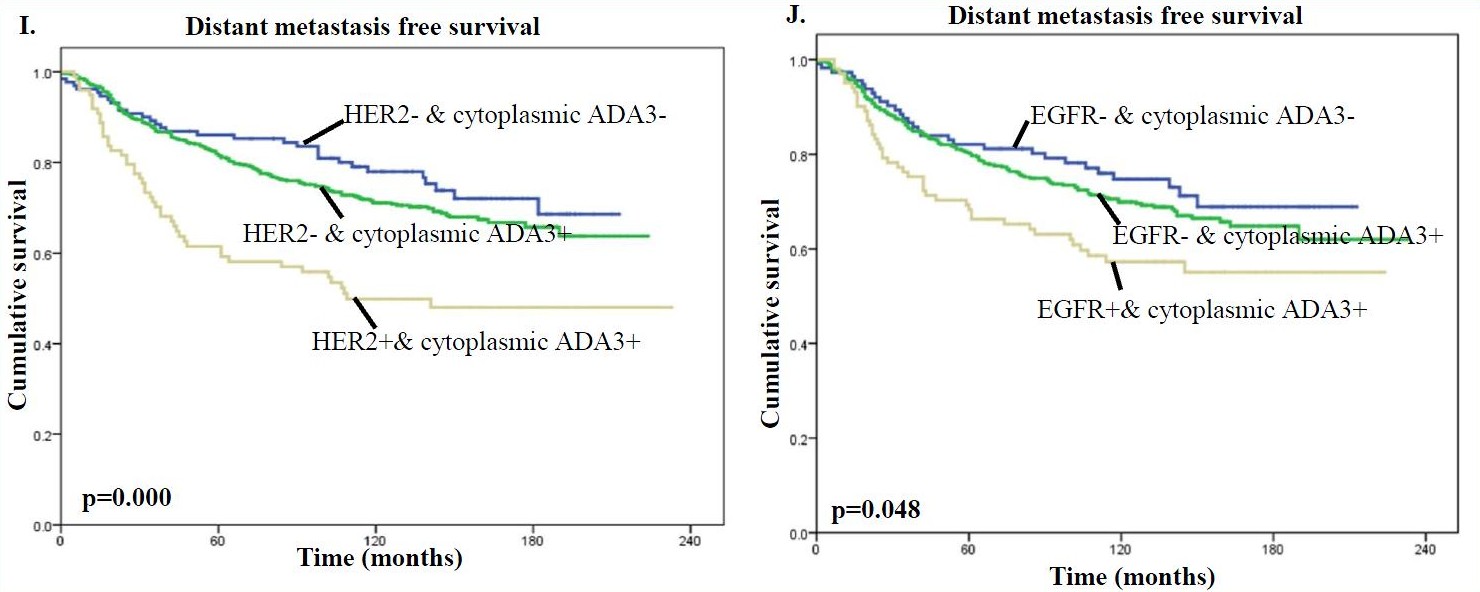
Therefore, when factoring in clinical outcome, the nuclear vs. cytoplasmic localization of ADA3 can be utilized as a prognostic marker together with commonly used markers such as ER, PR, HER2, and EGFR when evaluating patient prognosis and in developing treatment strategies.